Contact: Lynn Yarris (510) 486-5375, [email protected]
What does uncovering the true authorship of plays attributed to Shakespeare have to do with identifying our genetic ancestors or classifying new life forms? All involve the comparative analysis of long sets of data and all will benefit from a unique new analytical tool developed by researchers at Berkeley Lab.
Sung-Hou Kim, a chemist who holds a joint appointment with Berkeley Lab’s Physical Biosciences Division and UC Berkeley’s Chemistry Department, led the development of a technique called “feature frequency profiles” (FFP), that makes it possible to compare, classify, index and catalog just about any type of linear information that can be electronically stored. The kinds of information that can be analyzed with the FFP technique include nucleotide base and amino acid sequences, books, documents and possibly images. It could even prove to be the ultimate music organizer.
“I call our technique a tool for demographic phylogeny because it enables us to organize large sets of data into groups and find relationships among these groups,” says Kim. “The idea is to organize data sets into groups based on the frequency at which key features occur and then look for relationships. This is the reverse of what is usually done, where you find relationships in the data set then organize the data set into groups based on those relationships.”
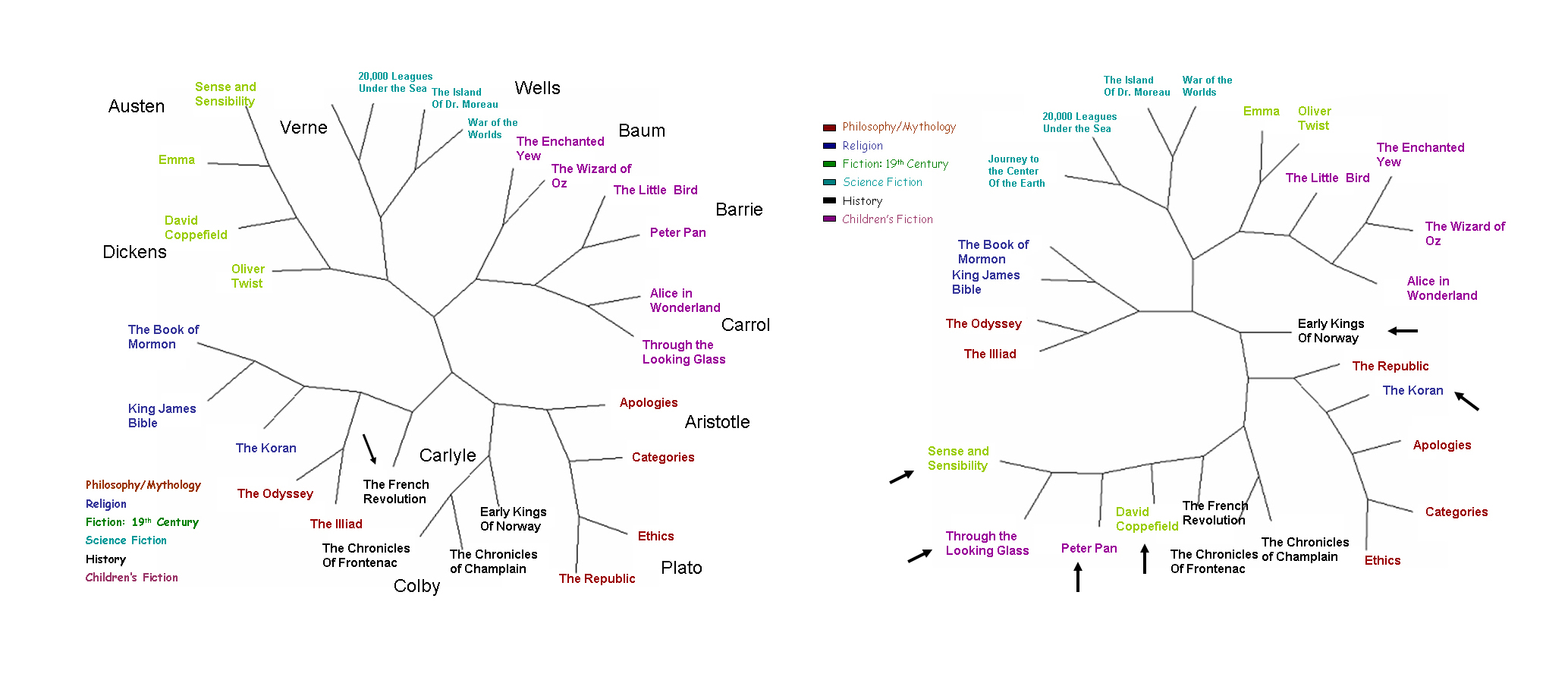
Using the FFP technique, Kim and his colleagues can create “family trees” that put into easy-to-see perspective the relationships between groups within a data set, whether those groups are books or genomes. The key is to identify the “optimal features” for profiling. For books, the optimal feature consisted of sequences of text about eight letters in length. For mammalian genomes, the optical feature consisted of sequences of nucleotide bases of about 18 base pairs in length. However, to keep their genomic computations manageable, Kim and his colleagues reduced the four-letter DNA alphabet (adenine, guanine, thymine and cytosine) to a two-letter alphabet by using R for the purine nucleic acids and Y for the pyrimidine nucleic acids). In a series of tests run on books and genomes, the FFP technique provided a more comprehensive and in some cases more accurate analysis over the standard analytical tools.
For example, Kim and his colleagues used the FFP technique to create a book tree composed of more than two dozen selected works under the categories of philosophy, mythology, religion, 19th Century fiction, science fiction and children’s fiction. Their FFP-based book tree correctly grouped all books by category and author including some, such as the Koran, that were misplaced in a book tree based on a standard word frequency profile analysis. In the case of the Koran, the FFP-based tree placed it in the religion category on the same branch as the King James Bible and the Book of Mormon, whereas the word frequency book tree grouped it in the philosophy category, on the same branch as Plato’s The Republic and Socrates’ The Apology.
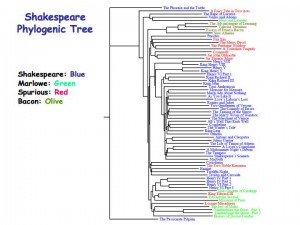
An FFP phylogeny tree of the works of William Shakespeare supports scholars who question the Bard’s authorship of Pericles. (Click to enlarge)
Kim and his colleagues later applied the FFP technique to a comparative analysis of the works of William Shakespeare, contemporaries such as Christopher Marlowe, plus several works from the Jacobean era that were once attributed to Shakespeare but whose authorships are now in question. The results cast new doubt on Shakespeare having been the author of the play Pericles, Prince of Tyre, and point to his authorship of the comedy Two Noble Kinsmen, for which in the past he has only received partial credit.
“I was stunned when I saw how well the technique worked with books,” Kim says.
The next step was the successful application of the technique to the whole genomes of mammals whose phylogenic tree is well established, then on to whole genomes of prokaryote organisms (bacteria and Archaea) and finally on to viruses, for which current comparative genomic analytic tools sometimes cannot be applied.
Collaborating with Kim on this project have been biophysicist Gregory Sims, statistical mathematician Se-Ran Jun and theoretical physicist Guohong Wu. Like Kim, they all hold joint appointments with Berkeley Lab and UC Berkeley.
Exploding Growth of Genome Data
Kim is an internationally recognized authority on protein structures and a pioneer in the field of structural genomics. In 2003, he unveiled a 3-D demographic map of the protein structure universe that for the first time made it possible to organize the structures of this vast assemblage of biological molecules (more than 50 billion known species and growing) into meaningful groups.
“Scientists studying the genomes of different organisms are facing similar problems to those studying protein structures, perhaps even more difficult,” Kim says. “Thousands of whole genomes have been or are in the process of being sequenced and we need to have an effective way of comparing and grouping them, and finding relationships among the groups. The FFP method can help us mine the function of gene-coding and non-coding nucleotide base sequences in the genome of a particular species, and can also give us a better understanding of how that species may have evolved, who its closest relatives are and other valuable information.”

Sung-Hou Kim, a world authority on proteins and structural genomics, holds a joint appointment with Berkeley Lab's Physical Biosciences Division and UC Berkeley's Chemistry Department. (Photo by Roy Kaltschmidt, Berkeley Lab Public Affairs)
Currently, comparative genomics studies are based either on measuring the similarities and differences between a set of selected genes in the coding regions of the genomes that are common to the species being compared, or on gene-profiles, in which the presence of certain genes in two or more species yields a similarity score. Species with a higher number of shared genes or similarity scores are presumed to be more closely related than those with a lower number. Both of these methods require an alignable set of common genes in the coding regions, which is not always the case, especially amongst the genomes of rapidly evolving species. Such “gene-centric” comparisons also suffer from an even greater limitation for comparing mammals and other high-order eukaryotes, as Kim explains.
“Coding sequences (exons) total only about one-percent of the entire human genome, with the rest made up of non-coding sequences (introns) whose functions are still largely unknown,” he says. “What is needed is an alignment-free method that can be used for comparing entire genomes or genomic regions that may be distantly related, have undergone significant rearrangement, or do not share a common set of genes. We also need a tool that can be used to analyze and compare nongenic regions of genomes as well.”
Kim began this quest by turning to the world of books, where comparative analytical tools are well established to ascertain authorship as well as to expose fraud or plagiarism. However, two problems became evident. First, current standard text analysis is based on the frequency at which different words appear, but genomic data consists of long strings of letters not words. Second, analysis based on the frequency of words does not provide local syntax – the relationship between adjoining words, a point that is critical in comparative genomics and turned out to be important in text comparisons as well.
A New Look at Books and Genomes
To overcome the limitations of current text comparison techniques, Kim and his colleagues first undertook an analysis of words in a Webster’s English dictionary and found that words with eight to nine letters were optimal for frequency profiling. This finding also proved true for all other books as well.
“Text features longer than eight or nine letters do not occur frequently enough for frequency profile comparisons, and text features shorter in length do not give us enough information to distinguish one book from another,” Kim says.
To apply their FFP technique to comparative analysis of books, they “delimiter-stripped” each book – meaning they stripped the text of all punctuation and spaces – then transformed the text into a single long string of letters. A “window” of eight letters in length was then advanced across this string one letter at a time, yielding a frequency profile of the features in which overlapping sequences of text reveal relationships between individual features. Comparing the feature frequency profile for each book analyzed produced astonishingly accurate trees that grouped books by author, genre or historical era.
“This enables us to capture the syntactical idiosyncrasies of specific authors as well as the unique vocabulary associated with a certain genres or subject matter,” says Kim. “When we saw the results of our book tree, we knew we were ready for genomes.”
Applying the FFP technique to whole genome sequences of mammals produced the exact same family tree as phylogenic trees constructed through traditional approaches based on genetic, morphological, anatomical and fossil data. Kim and his colleagues also used the FFP technique to investigate the existence of a “phylogenic signal” embedded within the non-coding regions of genomes.
“We found a high level of similarity between the phylogeny obtained from the non-coding FFP comparisons and the established gene-based consensus mammalian phylogeny,” Kim says. “It shows that evolutionary signals are imprinted in the entire genome and not just in the genes. We think the reason is that the sequence-changing mechanisms don’t know if they are changing in a coding or a non-coding region of a genome. In other words, mutations equally affect all parts of the genome, but may be selected or filtered out differently in non-coding versus coding sequences.”
In the final phase of their testing, Kim and his colleagues applied the FFP method to a total of 518 genomes, representing eukaryotes and prokaryotes, plus a couple of random genome sequences. For the prokaryotes – bacteria and Archaea, they used amino acid sequences, which are the building blocks of proteins instead of the base sequences used in mammals. This was done because unlike the genomes of mammals, the genomes of prokaryotes consist almost exclusively of sequences that code for proteins, which means that the “proteome” of these microorganisms (their entire complement of proteins) may hold the key to constructing accurate family trees for them.
“There is a lot of controversy surrounding prokaryotes with regards to which demographic group came first on the evolutionary tree,” Kim says. “We wanted to test whether our method could provide any new insight on this issue.”
Their results showed that the FFP method can be used to group bacteria and Archaea into separate domains, phyla and classes that are in general agreement with currently accepted grouping, but the evolutionary relationships among the groups came out different from those obtained
from traditional genetic and morphology studies. With their FFP technique, Kim and his colleagues were also able to classify microbes that had not been classified before. They also successfully used it to classify the genomes of several hundred viruses.
“No one has been able to figure out the evolutionary relationship between viral groups, but our FFP technique was able to suggest evolutionary relationships between some of these groups. We were very happy to see that,” says Kim.
Much work remains to be done with the FFP technique, Kim says, and some of the observations they have made thus far could eventually prove to be wrong. But the groundwork has been laid and with further improvements, the FFP technique could be expanded far beyond books and genomes into the world of music.
“We could really go wild,” Kim says, laughing, “and use it to organize all the books and documents, or even all the music ever written into useful demographic groupings.”
This work was funded by the National Institutes of Health and by a grant from the Korean Ministry of Education, Science and Technology. A paper describing the research has been published in the Proceedings of the National Academy of Sciences.
Additional Information:
For more about the research of Sung-Hou Kim, visit the Website at
http://chem.berkeley.edu/people/faculty/kim/kimgroup/index.html