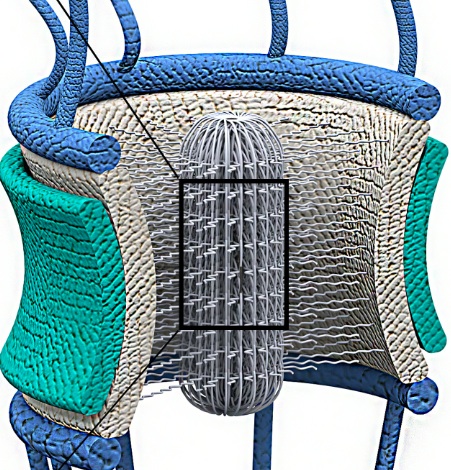
Scientists from the U.S. Department of Energy’s Lawrence Berkeley National Laboratory (Berkeley Lab) have uncovered new clues to how a molecular machine inside the cell acts as a gatekeeper, allowing some molecules to enter and exit the nucleus while keeping other molecules out.
The scientists studied the nuclear pore complex, which crosses the membrane that separates the nucleus from the rest of the cell. Amazingly, this nanomachine can handle the selective transport of more than one thousand molecules per second. The complex is critical to maintaining an organism’s health, including ours. It transports RNA and ribosomal proteins out of the nucleus, and allows only important molecules to enter. It’s also implicated in several diseases when it malfunctions, such as cancer, cardiac and infectious diseases, and autoimmune and nervous system disorders.
Despite decades of research, however, scientists don’t know what makes the nuclear pore complex both high throughput and highly selective. It’s well known the nanopore uses a family of proteins called FG Nups to transport molecules. These proteins are inside the pore and have floppy regions that briefly interact with molecules, allowing passage to some and blocking others. But it’s unclear how these proteins perform this job.
Now, as reported November 6 in the journal Scientific Reports, Berkeley Lab scientists found that the proteins’ cargo-transporting ability is likely due to specific features encoded in their sequences. Protein sequences are primarily composed of amino acids, and in FG Nups proteins, the scientists found different types of amino acids arranged just so within their sequences. This arrangement of amino acids appears to form a network inside the nuclear pore complex that is optimized for one purpose only: the extremely efficient transport of molecular cargo into and out of the nucleus.
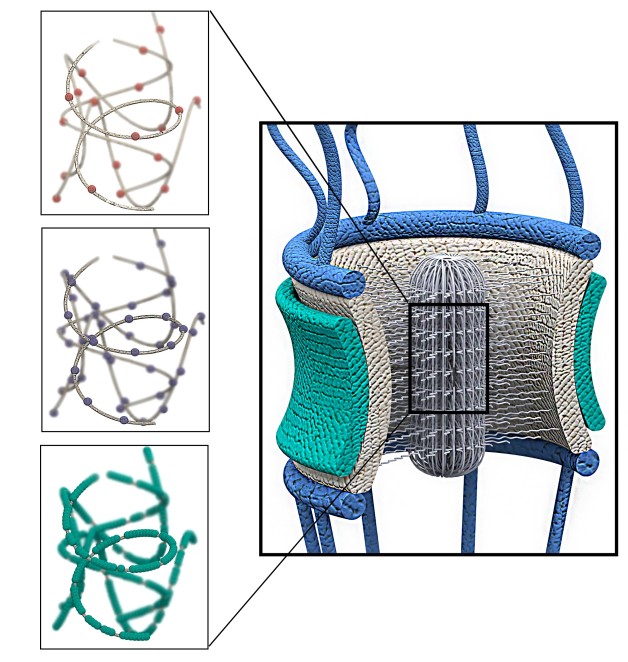
This schematic of a nuclear pore complex shows how different types of amino acids, meticulously arranged along the sequences of FG Nups proteins, form a network that is extremely efficient at transporting molecules. The three left boxes show “Like Charge Regions,” FG motifs, and polar amino acids, respectively. (Credit: Mohammad Mofrad, Berkeley Lab)
“Our results suggest a striking cooperation among the different types of amino acids that are meticulously positioned within the sequences of FG Nups,” says Mohammad Mofrad, a faculty scientist in Berkeley Lab’s Molecular Biophysics and Integrated Bioimaging Division and a professor of Bioengineering and Mechanical Engineering at UC Berkeley. Mofrad conducted the research with members of his UC Berkeley group.
They discovered this amino acid teamwork by analyzing the sequences of more than 1000 FG Nups proteins across 252 species.
“These evolutionary conserved features could well explain how FG Nups form networks to transport of cargo,” says Mofrad.
Specifically, the scientists identified polar amino acids, and a newly discovered sequence of charged amino acids, which they named “Like Charge Region,” as the likely players in forming FG Nups networks in nuclear protein complexes. They also found that positively charged “Like Charge Regions” prepare the pore for negatively charged cargos and regulate the size of the FG Nups network, while polar amino acids facilitate the formation of FG Nups-rich regions in the pore.
“This study is a step forward in building a more comprehensive understanding of nuclear pore complex function, which has numerous implications in health and disease. In addition, our results can set a basis for the design of nuclear pore complex-inspired nanopores for filtering and separation technologies,” Mofrad says.
This research was supported in part by the National Science Foundation.
###
Lawrence Berkeley National Laboratory addresses the world’s most urgent scientific challenges by advancing sustainable energy, protecting human health, creating new materials, and revealing the origin and fate of the universe. Founded in 1931, Berkeley Lab’s scientific expertise has been recognized with 13 Nobel prizes. The University of California manages Berkeley Lab for the U.S. Department of Energy’s Office of Science. For more, visit www.lbl.gov.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.