Researchers have shown that when parts of a genome known as enhancers are missing, the heart works abnormally, a finding that bolsters the importance of DNA segments once considered “junk” because they do not code for specific proteins.
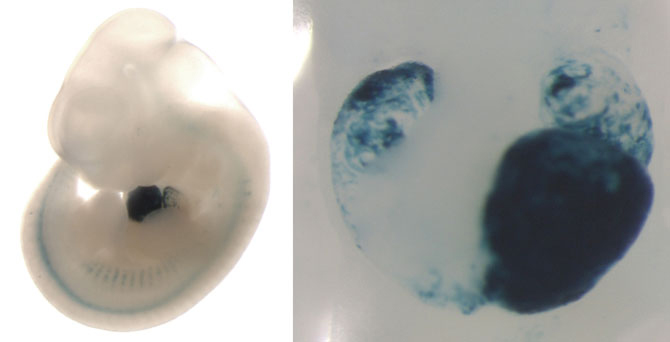
On the left, a mouse embryo showing enhancer activity (blue staining) in the developing heart. On the right, a closeup of this heart, showing that the enhancer is active in the left ventricle, left atrium, and right atrium of the heart. (Credit: Mammalian Functional Genomics Laboratory/Berkeley Lab)
The team, led by scientists at the Department of Energy’s Lawrence Berkeley National Laboratory (Berkeley Lab), examined the role of two heart enhancers in the mouse genome, showing that the loss of either one resulted in symptoms that resemble human cardiomyopathy, a disease in which the heart muscle often becomes enlarged or rigid. In humans, the disease often leads to heart failure.
The findings appear in a study published today in the journal Nature Communications.
In that same paper, the researchers provided a comprehensive genome-wide map of more than 80,000 enhancers considered relevant to human heart development and function. The two heart enhancers that they tested were the mouse equivalent of enhancers chosen from among that catalog.
“The cardiac changes that we observed in knockout mice lacking these enhancers highlight the role of noncoding sequences in processes that are important in human disease,” said study co-senior author Axel Visel, senior staff scientist and one of three lead researchers at the Mammalian Functional Genomics Laboratory, part of Berkeley Lab’s Environmental Genomics and Systems Biology (EGSB) Division. “Identifying and interpreting sequence changes affecting noncoding sequences is increasingly a challenge in human genetics. The genome-wide catalog of heart enhancers provided through this study will facilitate the interpretation of human genetic data sets.”
Study lead author Diane Dickel, project scientist, and co-senior author Len Pennacchio, senior staff scientist, both work with Visel at Berkeley Lab’s Mammalian Functional Genomics Laboratory.
DNA Dark Matter
When scientists sequenced the human genome, they discovered that less than 5 percent of our DNA were genes that actually coded for protein sequences. The biological functions of the noncoding portions of the genome were unclear.
Over the past fifteen years, however, there has been a growing appreciation for the importance of these noncoding regions, thanks in large part to the efforts of individual labs and, more recently, large international efforts such as the Encyclopedia of DNA Elements (ENCODE) project.
What became clear from this work is that there are many elements of the genome, including enhancers, that are involved in regulating gene expression, even though they do not encode for proteins directly.
This realization meant that there were vast sections of the genome that needed to be explored and understood. Dickel noted that there are about 20,000 genes in the mouse genome, and in many cases, scientists have a fairly good understanding of what will happen if any one of them is disabled. In contrast, there are 80,000 candidate heart enhancers in the human genome, and it is still unclear how important they are for human development.
“In genetic studies, the way you establish whether a gene is important is you delete it from the genome and see what happens,” said Dickel. “In many cases, there are genes that, if disabled, make it difficult for the organism to survive. For enhancers, it’s less known what the consequences are if they are damaged or missing. To use a car analogy, if we took the battery out of a car, it wouldn’t start. That’s a critical component. A missing or damaged enhancer could be essential like a battery, or more similar to a missing passenger seat in the car. It’s not as nice, but it’s still possible to drive the car.”
Mapping and Testing the Enhancers
To assess the function of heart enhancers, the researchers first compiled a single road map to guide them. They used results from a technology called ChIP-seq (chromatin immunoprecipitation sequencing) to identify the likely heart enhancers in the human genome.
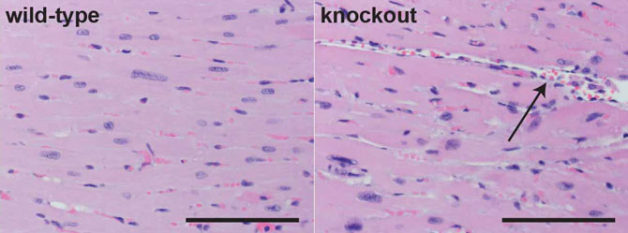
Heart tissue from a normal (wild-type) mouse, with cells stained pink and nuclei in blue, is shown on the left. Hearts from mice missing an important heart enhancer (knockout) exhibit disorganized cell growth, including large spaces between cells (arrow), as shown on the right. (Credit: UC Davis Comparative Pathology Laboratory)
The researchers say this map will become an important tool as advances in genomics usher in a new era of personalized medicine.
“This compendium of human heart enhancers will be a valuable resource for many disease researchers who have begun adopting whole genome sequencing of patients to look for disease-causing mutations in both the coding and noncoding portion of the genome,” said Dickel.
Using the map, the researchers picked two enhancers located near genes associated with human heart disease. They then determined their equivalent enhancers on the mouse genome and disabled them in mice.
They compared the mice with the disabled enhancers with control mice that had no mutation and saw very large changes in gene expression in the test mice.
Echocardiograms used to image the hearts from the two groups of mice confirmed that the heart tissue of mice with a disabled enhancer was pumping with less power than normal, consistent with the signs of human cardiomyopathy.
“Prior to this work, no study had looked at what happens to heart function as a result of knocking out the heart enhancers in the genome,” said Dickel. “What was surprising to me was that outwardly, the knockout mice seemed fine. If you just looked at them, you wouldn’t necessarily see anything wrong.”
With so many enhancers to test, the map could help scientists prioritize which ones to assess in animal studies and in disease research, the researchers said.
The National Institutes of Health provided funding for this research.
###
Lawrence Berkeley National Laboratory addresses the world’s most urgent scientific challenges by advancing sustainable energy, protecting human health, creating new materials, and revealing the origin and fate of the universe. Founded in 1931, Berkeley Lab’s scientific expertise has been recognized with 13 Nobel prizes. The University of California manages Berkeley Lab for the U.S. Department of Energy’s Office of Science. For more, visit www.lbl.gov.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.