It is said that a picture is worth a thousand words, but new images of structural fibers inside a cell may represent more than a million words from hundreds of research papers spanning the past three decades.

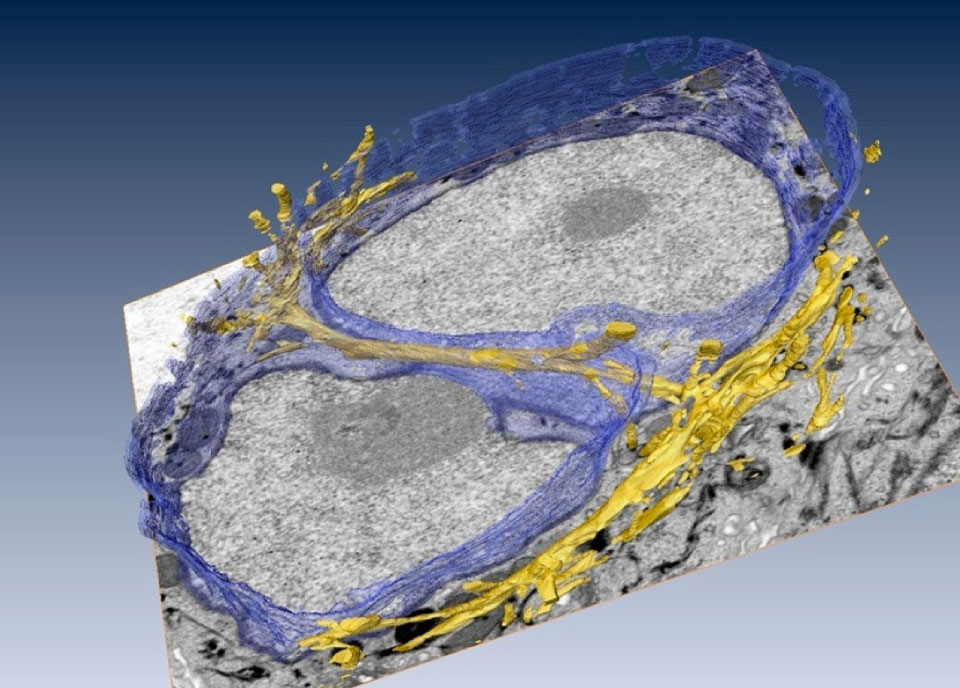
Rendering of the 3-D architecture of cytofilament bundles tunneling through a cell nucleus. (Credit: Manfred Auer/Berkeley Lab)
The images, obtained by scientists at the Department of Energy’s Lawrence Berkeley National Laboratory (Berkeley Lab), show thread-like cytofilaments reaching into and traversing a human breast cell’s chromatin-packed nucleus. It provides the first visual evidence of a physical link by which genes can receive mechanical cues from its microenvironment.
The images appear in a study featured on the cover of the Journal of Cell Science in a special issue on 3D Cell Biology published this month. The work leading up to the images began in the early 1980s when Berkeley Lab’s Mina Bissell proposed the idea that gene expression and cell fate were dependent on their physical surroundings called extracellular matrix.
“There are somewhere between 30-70 trillion cells in our bodies, all with the same DNA sequence, so I’ve been saying since 1981 that something other than the sequence of the genes had to allow a nose to be a nose and not an elbow,” said Bissell, Distinguished Scientist at Berkeley Lab’s Biological Systems and Engineering Division and co-corresponding author of this study. “When the shape changes, biology changes.”
The concept that phenotype is dominant over genotype was initially met with great skepticism, but it has since become accepted in the field. Before this, it was believed that the dominant signals dictating cellular function and form were controlled only by soluble small molecules such as hormones and growth factors, whereas extracellular matrix (ECM) molecules outside the cells were large insoluble proteins.
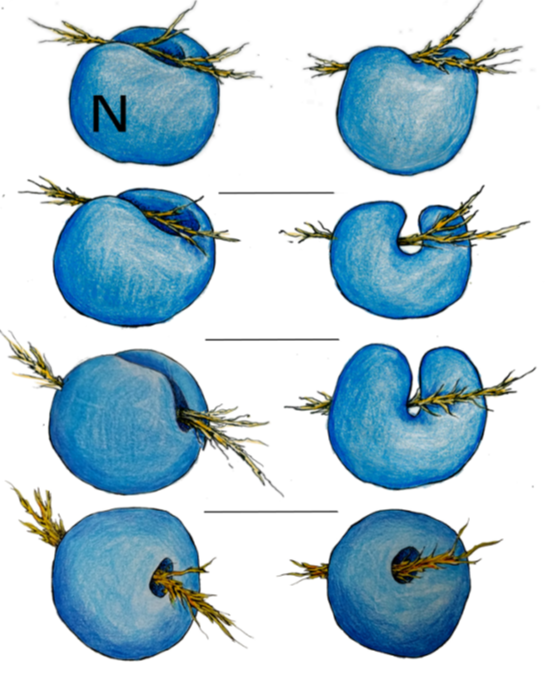
Artistic renderings of cytofilaments reaching deep into or tunneling through the cell nucleus. (Credit: Cerise Bennett/Berkeley Lab)
Evidence builds for mechanical influence
Hundreds of papers, including some 400 led or co-authored by Bissell, have since provided critical clues showing that signals from physical forces outside a cell could dramatically alter a cell’s function. By growing cells in a 3-D gel that includes extracellular matrix, researchers coaxed samples of breast cells from lactating mice to produce milk. This showed that cell function depended on having the proper 3-D growth environment.
“We knew the extracellular matrix was affecting gene expression, but it wasn’t understood until now that the cytoskeleton was actually able to connect inside the nucleus,” said Bissell. “Now we know there’s a direct connection to the nucleus. That’s what we’re showing here for the first time. This is absolutely novel.”
Bissell teamed up with Manfred Auer, head of the Cell and Tissue Imaging Department at Berkeley Lab’s Molecular Biophysics and Integrated Bioimaging Division and co-corresponding author of the study.
“It took advances in cryogenic sample preparation techniques and large-volume electron microscopy to come up with these images,” said Auer.
Also critical were developments in super-resolution imaging by study co-author Ke Xu, a Berkeley Lab faculty scientist and UC Berkeley assistant professor of chemistry. Specifically, Xu works with stochastic optical reconstruction microscopy, or STORM, to create sub-diffraction resolution images of cells.
Hundreds of millions of data points to get one image
“We combined a record-breaking six different imaging techniques and hundreds of millions of data points to obtain these images,” said Auer. “The integrative bioimaging approach involved three different optical light and three different electron microscopy imaging approaches, each with its own strengths. This new integration of imaging approaches is what allowed us to study something as complex as this cytofilaments system.”
With the clarity provided by the super-resolution imaging, the researchers could show that the cytoskeleton coalesced with SUN proteins, a type of protein involved in connections between the donut-shaped nucleus and cell cytoplasm.
“This study establishes for the first time the long-postulated mechanical link between the cell’s nucleus to adhesion complexes that allow communication with the surrounding extracellular matrix and other cells,” said Auer.
What had been previously seen through other imaging techniques were telltale cytoskeletal tracks going through the cytoplasm of the cell, but it took this high-powered integrated bioimaging used in the study to reveal deep invaginations into and through the cell nucleus. The invaginations contained cytofilaments anchored at the nuclear membrane, thus providing a macromolecular highway allowing cables of cytofibers, which are known to interact with the extracellular matrix, to travel from the outside of the cell to its nucleus.
“The reason we’re excited is that it explains a whole lot of literature of how force and tension could be playing a role together with biochemical signals to bring about huge changes in a cell,” said Bissell.
Co-lead authors of the study are Berkeley graduate students Danielle Jorgens, Jamie Inman, and Michal Wojcik; and postdoctoral fellow Claire Robertson. Jorgens is now a research assistant professor of biomedical engineering at Oregon Health & Science University, and Inman is a postdoctoral fellow in the Bissell Lab.
Funding for this research was provided by the National Institutes of Health, the Department of Defense’s Breast Cancer Research Program and the Breast Cancer Research Foundation in New York. The National Science Foundation helped support the foundation of this work.
###
Lawrence Berkeley National Laboratory addresses the world’s most urgent scientific challenges by advancing sustainable energy, protecting human health, creating new materials, and revealing the origin and fate of the universe. Founded in 1931, Berkeley Lab’s scientific expertise has been recognized with 13 Nobel Prizes. The University of California manages Berkeley Lab for the U.S. Department of Energy’s Office of Science. For more, visit www.lbl.gov.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.