An international team of scientists, including a researcher from Berkeley Lab, has characterized the genome of Biomphalaria glabrata, a freshwater snail that transmits a parasitic worm responsible for the infectious disease schistosomiasis, also known as snail fever.
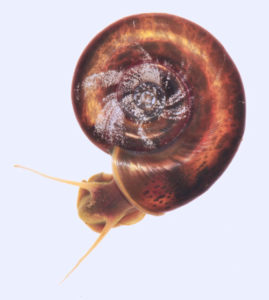
Biomphalaria glabrata, a freshwater snail that carries a parasitic flatworm responsible for the infectious disease schistosomiasis. (Credit: Tom Kennedy and Coen Adema/UNM)
The genome analysis could help researchers disrupt the life cycle of Schistosoma mansoni, a parasitic flatworm transmitted to humans through contact with contaminated freshwater.
The snails carry the worm eggs, which hatch in freshwater and can penetrate the skin of humans. Inside the human host, the worm can lead to anemia, abdominal bleeding, and enlargement of the liver and lungs, among other complications.
The study was led by Coen Adema at the University of New Mexico. Berkeley Lab’s Monica Munoz-Torres led the biocuration of experimental data and literature related to the structure and localization of the snail’s genes.
“We provided guideposts that helped identify candidates for the genes in this study,” said Munoz-Torres, a bioinformatics scientist in the Environmental Genomics and Systems Biology Division.
To do this, she relied upon an open-source genome annotation editor called Apollo. The free, web-based software was first released five years ago by Berkeley Lab scientists to support the community-based curation of genomes, and is now used by thousands of scientists all over the world.
The World Health Organization notes that more than 240 million people worldwide, mainly in tropical and subtropical climates, need preventive treatment for schistosomiasis.
To read the Nature Communications study, click here.
