Rice is a staple food for over half of the world’s population and a model for studies of candidate bioenergy grasses such as sorghum, switchgrass, and Miscanthus. To optimize crops for biofuel production, scientists are seeking to identify genes that control key traits such as yield, resistance to disease, and water use efficiency.
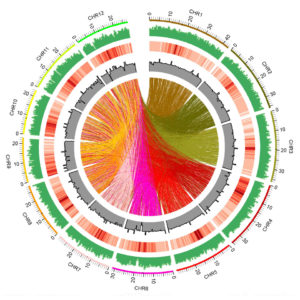
Genome-wide distribution of fast-neutron-induced mutations in the Kitaake rice mutant population (green). Even distribution of mutations is important to achieve saturation of the genome. Colored lines (center) represent translocations of DNA fragments from one chromosome to another. (Credit: Guotian Li and Rashmi Jain/Berkeley Lab)
Populations of mutant plants, each one having one or more genes altered, are an important tool for elucidating gene function. With whole-genome sequencing at the single nucleotide level, researchers can infer the functions of the genes by observing the gain or loss of particular traits. But the utility of existing rice mutant collections has been limited by several factors, including the cultivars’ relatively long six-month life cycle and the lack of sequence information for most of the mutant lines.
In a paper published in The Plant Cell, a team led by Pamela Ronald, a professor in the Genome Center and the Department of Plant Pathology at UC Davis and director of Grass Genetics at the Department of Energy’s (DOE’s) Joint BioEnergy Institute (JBEI), with collaborators from UC Davis and the DOE Joint Genome Institute (JGI), reported the first whole-genome sequenced fast-neutron induced mutant population of Kitaake, a model rice variety with a short life cycle.
Kitaake (Oryza sativa L. ssp. japonica) completes its life cycle in just nine weeks and is not sensitive to photoperiod changes. This novel collection will accelerate functional genetic research in rice and other monocots, a type of flowering plant species that includes grasses.
“Some of the most popular rice varieties people use right now only have two generations per year. Kitaake has up to four, which really speeds up functional genomics work,” said Guotian Li, a project scientist at Lawrence Berkeley National Laboratory (Berkeley Lab) and deputy director of Grass Genetics at JBEI.
In a previously published pilot study, Li, Mawsheng Chern, and Rashmi Jain, co-first authors on The Plant Cell paper, demonstrated that fast-neutron irradiation produced abundant and diverse mutations in Kitaake, including single base substitutions, deletions, insertions, inversions, translocations, and duplications. Other techniques that have been used to generate rice mutant populations, such as the insertion of gene and chromosome segments and the use of gene editing tools like CRISPR-Cas9, generally produce a single type of mutation, Li noted.
“Fast-neutron irradiation causes different types of mutations and gives different alleles of genes so we really can get something that’s not achievable from other collections,” he said.
Whole-genome sequencing of this mutant population – 1,504 lines in total with 45-fold coverage – allowed the researchers to pinpoint each mutation at a single-nucleotide resolution. They identified 91,513 mutations affecting 32,307 genes, 58 percent of all genes in the roughly 389-megabase rice genome. A high proportion of these were loss-of-function mutations.
Using this mutant collection, the Grass Genetics group identified an inversion affecting a single gene as the causative mutation for the short-grain phenotype in one mutant line with a population containing just 50 plants. In contrast, researchers needed more than 16,000 plants to identify the same gene using the conventional approach.
“This comparison clearly demonstrates the power of the sequenced mutant population for rapid genetic analysis,” said Ronald.
This high-density, high-resolution catalog of mutations, developed with JGI’s help, provides researchers opportunities to discover novel genes and functional elements controlling diverse biological pathways. To facilitate open access to this resource, the Grass Genetics group has established a web portal called KitBase, which allows users to find information related to the mutant collection, including sequence, mutation and phenotypic data for each rice line. Additional information about the database can be found through JGI.
Additional Berkeley Lab scientists who contributed to this work include co-first authors Rashmi Jain and Mawsheng Chern; Tong Wei and Deling Ruan, both affiliated with JBEI’s Feedstocks Division and with Berkeley Lab’s Environmental Genomics and Systems Biology Division; Nikki Pham and Kyle Jones of JBEI’s Feedstocks Division; and Joel Martin, Wendy Schackwitz, Anna Lipzen, Diane Bauer, Yi Peng, and Kerrie Barry of the JGI.
Support for the research at JBEI, a DOE Bioenergy Research Center, and JGI, a DOE Office of Science User Facility, was provided by DOE’s Office of Science. Additional support was provide by the National Institutes of Health and the National Science Foundation.
###
Lawrence Berkeley National Laboratory addresses the world’s most urgent scientific challenges by advancing sustainable energy, protecting human health, creating new materials, and revealing the origin and fate of the universe. Founded in 1931, Berkeley Lab’s scientific expertise has been recognized with 13 Nobel Prizes. The University of California manages Berkeley Lab for the U.S. Department of Energy’s Office of Science. For more, visit www.lbl.gov.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.
