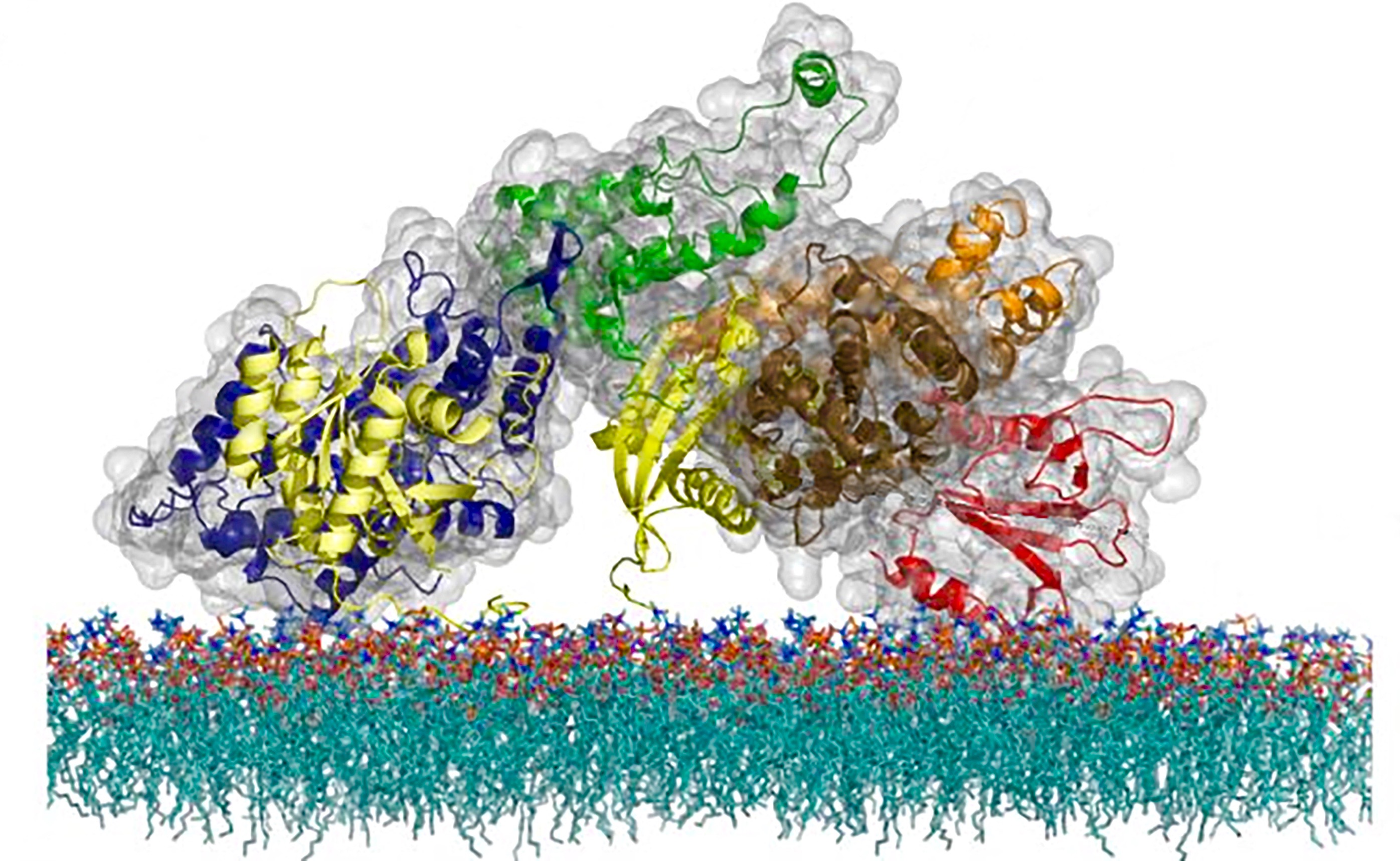
The atomic structure of the SOS protein, a cell messaging molecule that uses a unique timing mechanism to regulate activation of a critical immune system pathway. (Credit: Steven Alvarez/Berkeley Lab)
For the first time ever, scientists have imaged the process by which an individual immune system molecule is switched on in response to a signal from the environment, leading to the critical discovery that the activation process involves hundreds of proteins suddenly coming together to form a linked network through a process known as a phase transition.
The new work, described in a paper recently published in Science, provides an enormous leap forward in our understanding of how the immune system is finely tuned to detect even a single virus molecule amidst a sea of millions of other molecules, allowing us to quickly recover from viral infections such as the flu. By learning how these particular proteins work, scientists will also have a better grasp on why their activity sometimes goes awry – events that can lead to autoimmune diseases, such as diabetes or rheumatoid arthritis – and may provide unique insights into how to direct a cancer patient’s own immune system to cure cancer.
“This is something that happens inside a living cell during the process of the cell making a decision – signal transduction is what we call it – and it’s how cells ‘think’ with chemical reactions,” said study leader Jay Groves, a faculty chemist in the Biosciences Area of the Department of Energy’s Lawrence Berkeley National Laboratory (Berkeley Lab). “In the field of biology as a whole, the idea of a protein condensation phase transition has gained a lot of attention recently. Many groups all over the world study these phenomena, but until now, nobody knew how or why the cell is using them.
“Our paper is, I believe, the first to directly test and confirm how a phase transition can regulate signaling,” Groves said. “And the big discovery is that it’s a molecular timing mechanism. The cell is using time to distinguish a genuine receptor stimulation from background chemical noise.”
Studying an essential cell messenger
The team’s revelation came about as part of the ongoing research by the Groves lab on the physical mechanisms of T cell signaling and the Ras protein. Found in all eukaryotic cells in multiple variations, Ras wears many hats, including acting as a regulator for cell growth, division, and death. T cells, the immune system cells that detect foreign and potentially harmful molecules, use Ras as an on-off switch for the intruder alert pathway that launches a protective response. The T cell’s ability to distinguish a real external signal – when a foreign molecule binds with the aptly named T-cell receptor (TCR) on the cell surface – from inadvertent contact with nearby proteins is critical for a functioning immune system. If a T cell accidentally reacts to one of our own molecules, then an autoimmune disease can develop. At the same time, if a T cell loses its sensitivity, then viruses will be able to grow unchecked and cancerous cells won’t get cleared from the body.
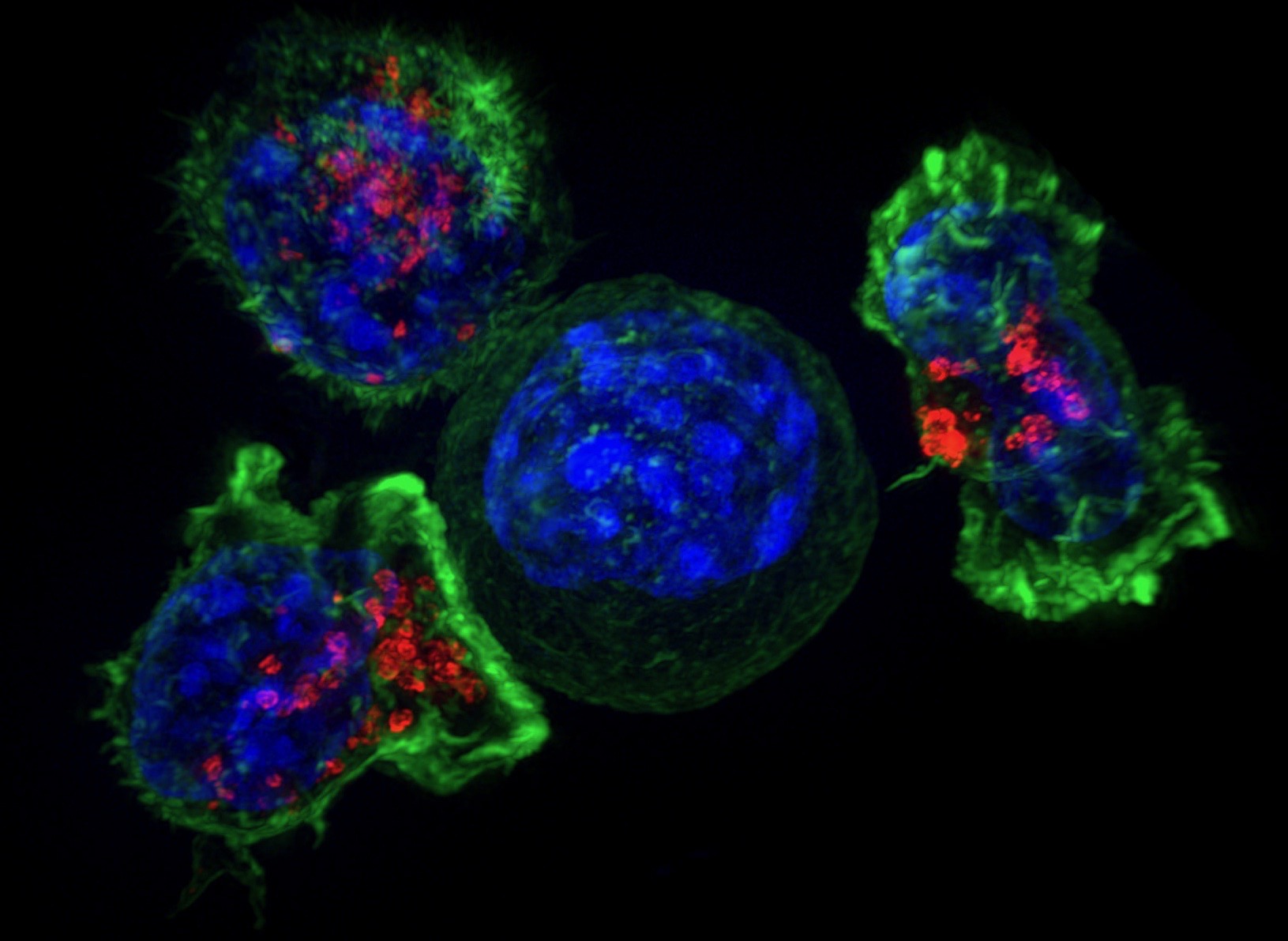
Killer T cells (green and red) surround a cancer cell (blue, center). Killer T cells are a type of immune cells that detect and remove unhealthy cells, including cancer cells and virus-infected cells. (Credit: NICHD/J. Lippincott-Schwartz)
Due to the wide implications for human health, scientists have long wondered how the cells regulate their signals in order to achieve this balance. Past research had revealed that a T cell’s Ras proteins don’t interact directly with cell receptors. Instead, receptors send the “on” signal to internal intermediary proteins, including a key group of three proteins, known as LAT, Grb2, and SOS, that ultimately pass the signal to Ras. Prior to this study, scientists knew that this molecular trio could link together in a phase transition process, but no one knew what the phase transition was doing. And until recently, it was impossible to figure out, because there was no technology that allowed scientists to directly monitor the activity from individual molecules in complex cell membrane systems.
The research team removed this hurdle by inventing an approach based on supported membrane microarrays, a technology the team has been developing for many years that uses scaffolds made of nanofabricated structures to hold cell membranes.
Phase transitions at work
In the current study, the scientists used microscopy to watch the moment that a T cell receptor on a supported membrane microarray asked a single SOS molecule to activate. Instead of responding right away, SOS waited for 10 to 30 seconds before turning into its active state. If the nearby LAT and Grb2 molecules underwent the phase transition with SOS, and condensed into their assembled state, they could hold SOS on the membrane long enough for SOS to activate. Without the phase transition, the long delay in the SOS molecule would prevent it from activating before it left the receptor.
“It’s like the protein has a delay built into it,” explained Groves. “It needs the phase transition combined with sustained signaling, and only then will it turn on.”
Though this research was specific to T-cell signaling, Groves and his colleagues believe that similar phase transition timing mechanisms are likely involved in a variety of other cell responses. Now that they established a proven experimental technique for observing the molecular activation of such processes, the team hopes to untangle more longstanding mysteries of how cells perform so many complex tasks.
In addition to his affiliation with Berkeley Lab’s Biosciences Area, Groves is a professor of chemistry at UC Berkeley. The other authors on this paper were William Huang, Steven Alvarez, Yasushi Kondo, Young Kwang Lee, Jean Chung, Hin Yue Monatrice Lam, Kabir Biswas, and John Kuriyan. This work is part of a larger international collaboration under the Center for Geometrically Engineered Cellular Systems, involving researchers from Berkeley Lab, UC Berkeley, UC San Francisco, and the University of Copenhagen. The research was funded by grants from the National Institutes of Health’s National Cancer Institute and the Novo Nordisk Foundation. The supported membrane microarray technology platform was developed under programs funded by the Department of Energy’s Office of Science.
# # #
Founded in 1931 on the belief that the biggest scientific challenges are best addressed by teams, Lawrence Berkeley National Laboratory and its scientists have been recognized with 13 Nobel Prizes. Today, Berkeley Lab researchers develop sustainable energy and environmental solutions, create useful new materials, advance the frontiers of computing, and probe the mysteries of life, matter, and the universe. Scientists from around the world rely on the Lab’s facilities for their own discovery science. Berkeley Lab is a multiprogram national laboratory, managed by the University of California for the U.S. Department of Energy’s Office of Science.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.