
Researcher Christine Shulse tends to Arabidopsis plants in a lab at the DOE Joint Genome Institute (JGI). (Photo credit: Marilyn Chung/Berkeley Lab)
An open-source RNA analysis platform has been successfully used on plant cells for the first time – an advance that could herald a new era of fundamental research and bolster efforts to engineer more efficient food and biofuel crop plants.
The technology, called Drop-seq, is a popular method for measuring the RNA present in individual cells, allowing scientists to see what genes are being expressed and how this relates to the specific functions of different cell types. Developed at Harvard Medical School in 2015, the freely shared protocol had previously only been used in animal cells.
“This is really important in understanding plant biology,” said lead researcher Diane Dickel, a scientist at the Department of Energy’s Lawrence Berkeley National Lab (Berkeley Lab). “Like humans and mice, plants have multiple cell and tissue types within them. But learning about plants on a cellular level is a little bit harder because, unlike animals, plants have cell walls, which make it hard to open the cells up for genetic study.”
For many of the genes in plants, we have little to no understanding of what they actually do, Dickel explained. “But by knowing exactly what cell type or developmental stage a specific gene is expressed in, we can start getting a toehold into its function. In our study, we showed that Drop-seq can help us do this.”
“We also showed that you can use these technologies to understand how plants respond to different environmental conditions at a cellular level – something many plant biologists at Berkeley Lab are interested in because being able to grow crops under poor environmental conditions, such as drought, is essential for our continued production of food and biofuel resources,” she said.
Dickel, who studies mammalian genomics in Berkeley Lab’s Environmental Genomics and Systems Biology Division, has been using Drop-seq on animal cells for several years. An immediate fan of the platform’s ease of use and efficacy, she soon began speaking to her colleagues working on plants about trying to use it on plant cells.
However, some were skeptical that such a project would work as easily. First off, to run plant cells through a single-cell RNA-seq analysis, they must be protoplasted – meaning they must be stripped of their cell walls using a cocktail of enzymes. This process is not easy because cells from different species and even different parts of the same plant require unique enzyme cocktails.
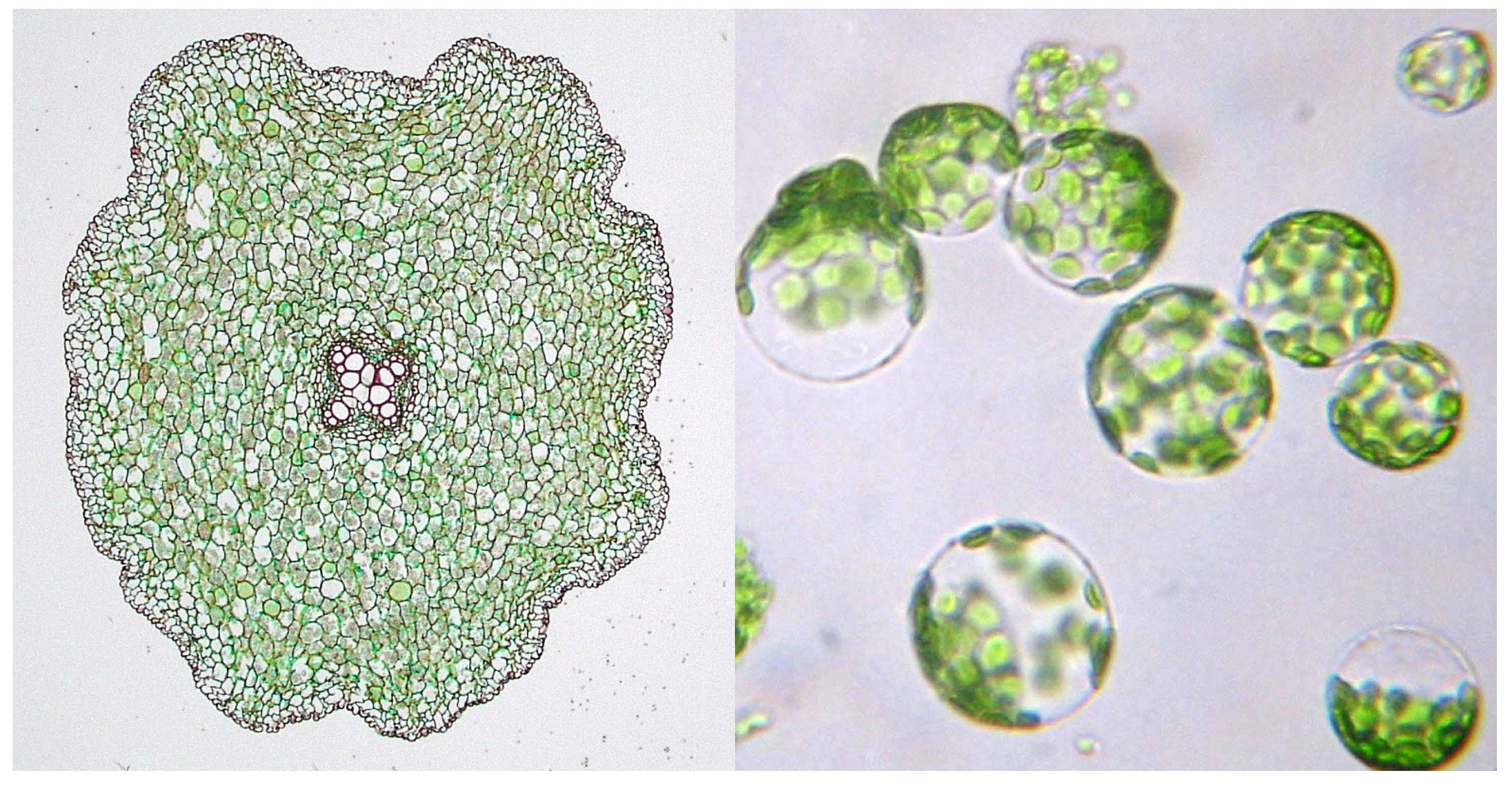
Microscope images of flowering plant root cells in their natural state (left) and after protoplasting (right). (Image credit: Berkshire Community College Bioscience Image Library and Department of Biological Sciences, Louisiana State University)
Secondly, some plant biologists have expressed concern that cells are altered too significantly by protoplasting to provide insight into normal functioning. And finally, some plant cells are simply too big to be put through existing single-cell RNA-seq platforms. These technologies, which emerged in the past five years, allow scientists to assess the RNA inside thousands of cells per run; previous approaches could only analyze dozens to hundreds of cells at a time.
Undeterred by these challenges, Dickel and her colleagues at the DOE Joint Genome Institute (JGI) teamed up with researchers from UC Davis who had perfected a protoplasting technique for root tissue from Arabidopsis thaliana (mouse-ear cress), a species of small flowering weed that serves as a plant model organism.
After preparing samples of more than 12,000 Arabidopsis root cells, the group was thrilled when the Drop-seq process went smoother than expected. Their full results were published this week in Cell Reports.
“When we would pitch the idea to do this in plants, people would bring up a list of reasons why it wouldn’t work,” said Dickel. “And we would say, ‘ok, but let’s just try it and see if it works’. And then it really worked. We were honestly surprised how straightforward it actually ended up being.”
The open-source nature of the Drop-seq technology was critical for this project’s success, according to co-author Benjamin Cole, a plant genomics scientist at JGI. Because Drop-seq is inexpensive and uses easy-to-assemble components, it gave the researchers a low-risk, low-cost means to experiment. Already, a wave of interest is building. In the time leading up to their paper’s publication, Dickel and her colleagues began receiving requests – from other scientists at Berkeley Lab, JGI, and beyond – for advice on how to adapt the platform for other projects.
“When I first spoke to Diane about trying Drop-seq in plants I recognized the huge potential, but I thought it would be difficult to separate plant cells rapidly enough to get useful data,” said John Vogel, lead scientist of plant functional genomics at JGI. “I was shocked to see how well it worked and how much they were able to learn from their initial experiment. This technique is going to be a game changer for plant biologists because it allows us to explore gene expression without grinding up whole plant organs, and the results aren’t muddled by signals from the few most common cell types.”
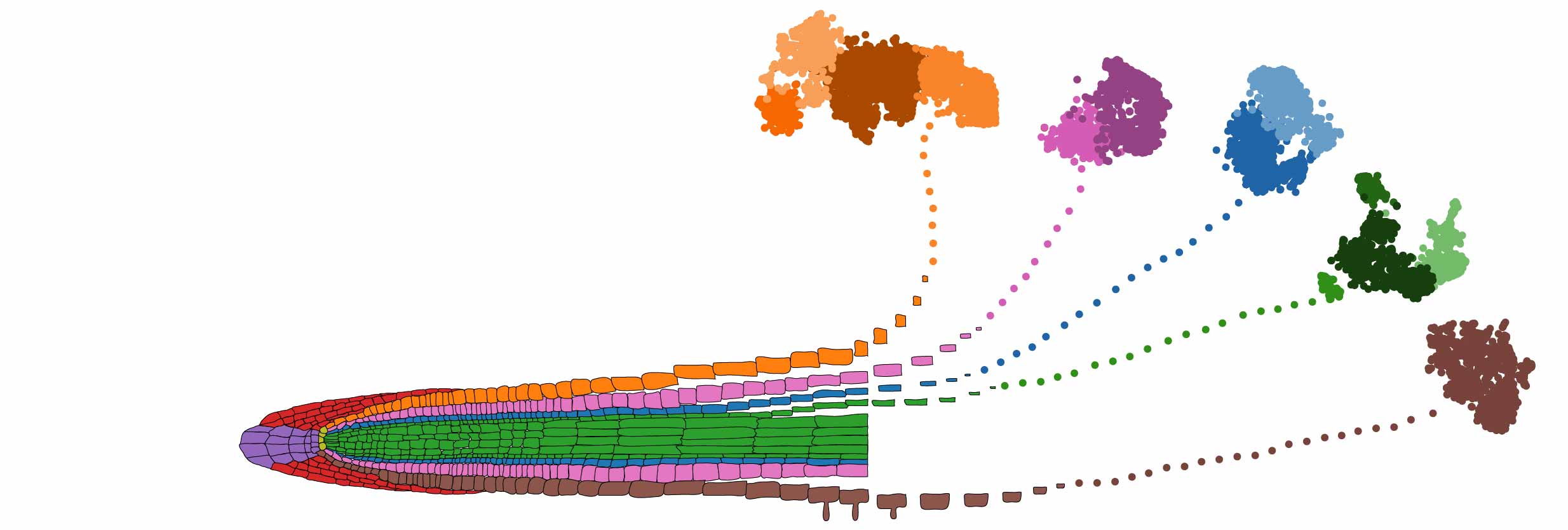
A cartoon diagram of the 17 different root cell types profiled using the Drop-seq protocol. (Image credit: Diane Dickel/Berkeley Lab)
The authors anticipate that the platform, and other similar RNA-seq technologies, will eventually become routine in plant investigations. The main hurdle, Dickel noted, will be developing protoplasting methods for each project’s plant of interest.
“Part of Berkeley Lab’s mission is to better understand how plants respond to changing environmental conditions, and how we can apply this understanding to best utilize plants for bioenergy,” noted first author Christine Shulse, who is currently a JGI affiliate. “In this work, we generated a map of gene expression in individual cell types from one plant species under two environmental conditions, which is an important first step.”
JGI is a DOE Office of Science user facility that was originally founded to advance the landmark Human Genome Project. After helping set the stage for a new era of medical and developmental science, JGI turned its focus to investigating how plants and microbes can provide solutions to pressing energy and environmental challenges.
This research was funded by the Laboratory Directed Research and Development (LDRD) program. The other authors were Doina Ciobanu, Junyan Lin, Yuko Yoshinaga, Mona Gouran, Gina Turco, Yiwen Zhu, Ronan O’Malley, and Siobhan Brady.
# # #
Founded in 1931 on the belief that the biggest scientific challenges are best addressed by teams, Lawrence Berkeley National Laboratory and its scientists have been recognized with 13 Nobel Prizes. Today, Berkeley Lab researchers develop sustainable energy and environmental solutions, create useful new materials, advance the frontiers of computing, and probe the mysteries of life, matter, and the universe. Scientists from around the world rely on the Lab’s facilities for their own discovery science. Berkeley Lab is a multiprogram national laboratory, managed by the University of California for the U.S. Department of Energy’s Office of Science.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.