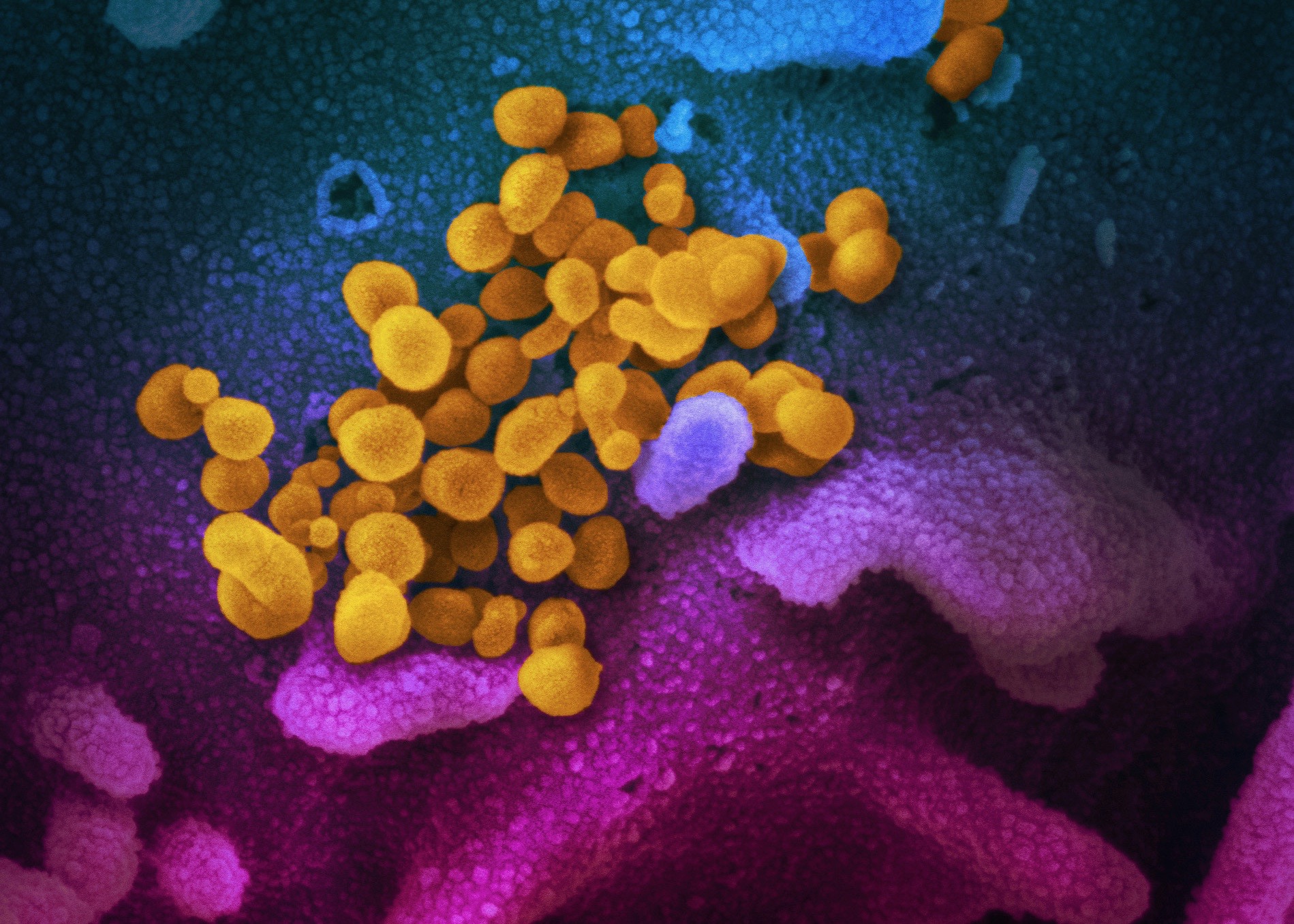
This scanning electron microscope image shows SARS-CoV-2 emerging from the surface of cells (blue/pink) cultured in the lab. (Credit: National Institute of Allergy and Infectious Diseases-Rocky Mountain Laboratories, NIH)
As the COVID-19 pandemic enters its second year, scientists are still working to understand how the SARS-CoV-2 strain evolved, and how it became so much more dangerous than other coronaviruses, which humans have been living alongside for millennia.
Virologists and epidemiologists worldwide have speculated for months that a protein called ORF8 likely holds the answer, and a recent study by Berkeley Lab scientists has helped confirm this hypothesis.
In a paper published in mBio, lead author Russell Neches and his colleagues show that ORF8 evolved from another coronavirus protein called ORF7a, and that both proteins have folds similar to that of a human antibody. This finding helps to explain how the virus avoids immune detection and is able to escalate into a severe infection in some hosts.
“By exploring the structural and functional characteristics of ORF8, and using supercomputers to look at the genomes of over 200,000 viruses, we discovered a striking and highly unusual evolutionary strategy,” said co-author Nikos Kyrpides, a computational biologist at the DOE Joint Genome Institute (JGI). “Amazingly, it seems that within the SARS clade, the gene encoding ORF7a is used as a ‘template’ gene, remaining stable, with a duplicate copy of this gene evolving to a point almost beyond recognition.” SARS-CoV-2 arose and exploded into a pandemic when a SARS strain’s duplicate ORF7a gene happened to mutate leading to a new protein (which we now call ORF8) that gave it the ability to interfere with immune cells.
According to the team, a similar event occurred in the SARS-CoV strain that caused the SARS epidemic in the early 2000s. In that instance, a copy of the ORF7a gene split into two, resulting in ORF8a and ORF8b proteins.
Christos Ouzounis, senior author of the study and a JGI affiliate scientist, noted that the connection between ORF8 and ORF7a was initially quite difficult to make, due to how little was known about this set of genes and their encoded proteins compared with the existing knowledge about surface proteins (such as the infamous spike protein), and because ORF8 and ORF7a currently seem wildly different. ORF7a is highly stable, mutation resistant protein that interacts with very few mammalian host proteins, whereas ORF8 is encoded by the most mutation prone gene in the viral genome, and is now known to be involved in dozens of interactions in the human body.
“Our findings – and their confirmation by parallel sequence and structure studies – reveal ORF8 to be an evolutionary hotspot in the SARS lineage. The lack of knowledge about the role of these genes has diverted attention to the more well-understood genes, but we now know more about this gene and hopefully it will receive more attention from the community,” said Kyrpides.
This work was supported by the ExaBiome Project, a Berkeley Lab-led collaboration that develops supercomputing tools for microbiome analysis. JGI is an Office of Science user facility.