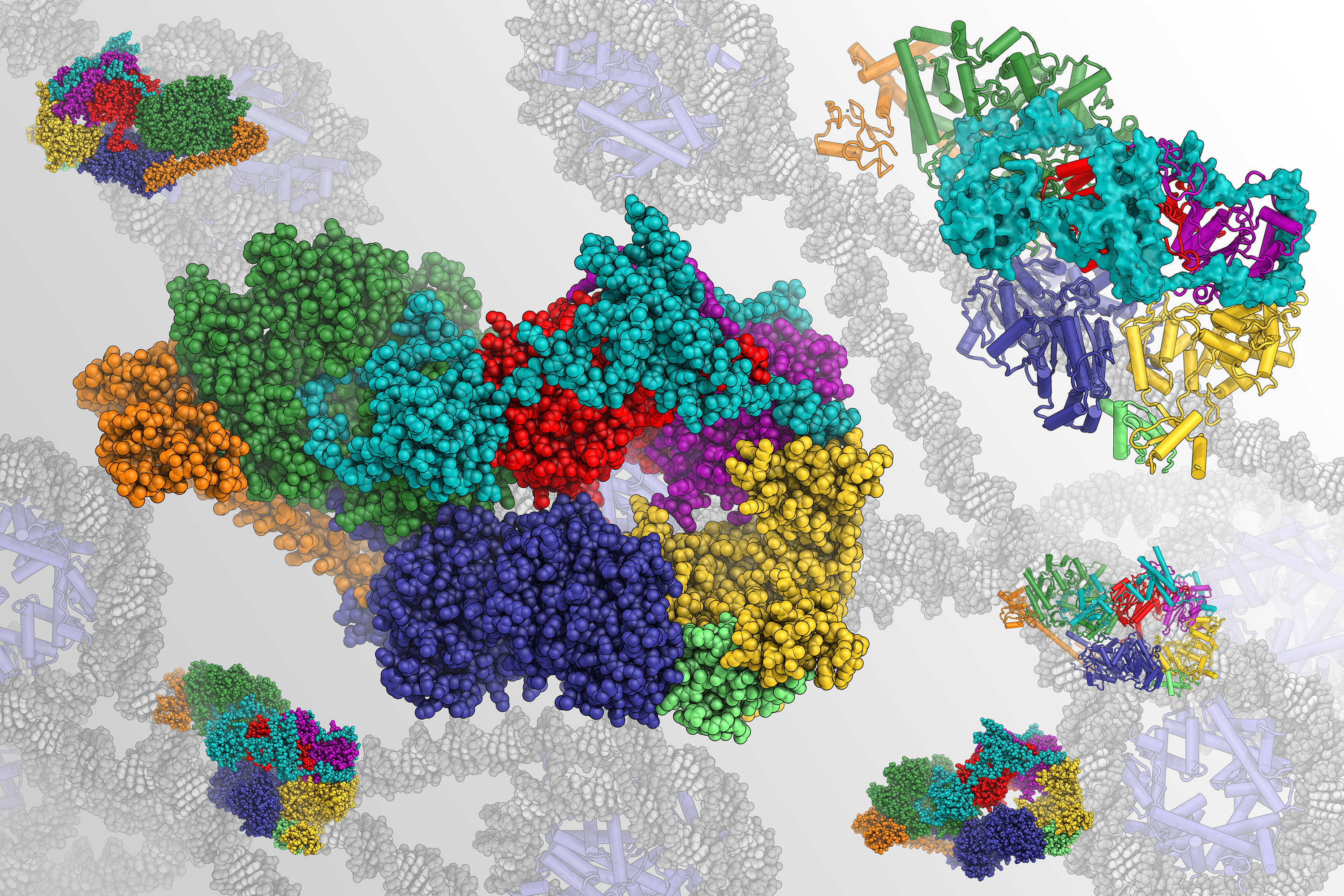
This video shows an animated structural model of TFIIH, a large protein assembly that helps repair and read our genome. Berkeley Lab scientists used an advanced cryo-electron microscope to generate the first-ever atomic map of TFIIH’s structure. (Credit: Basil Greber/Berkeley Lab)
Inside every cell in your body are molecular machines that help package, read, and repair DNA. These protein assemblies are essential to survival, yet we know little about how they function because, until recently, it was impossible to accurately describe their structure.
Too small to see with a light microscope and too intricate and scarce to image using traditional X-ray-based methods, molecules of this nature are finally being mapped out at the atomic level thanks to next-generation cryo-electron microscopy (cryo-EM). As reported in the journal eLife, a team of scientists from the Department of Energy’s Lawrence Berkeley National Laboratory (Berkeley Lab) and UC Berkeley used an advanced cryo-EM facility to construct the first complete atomic blueprint of an intriguing, multi-part machine called human transcription initiation factor IIH, or TFIIH.
Mutations in TFIIH are the cause of several inherited diseases that are characterized by premature aging and predisposition to cancer. Past research has also demonstrated that the TFIIH assembly could be a target for anti-cancer drug treatments.
The new work has already resolved several long-standing questions about how TFIIH works and is expected to advance research into how abnormal activity can lead to disease. TFIIH has been a subject of keen scientific interest since it was discovered in the late 80s.
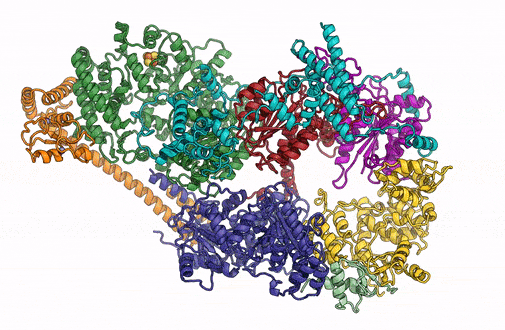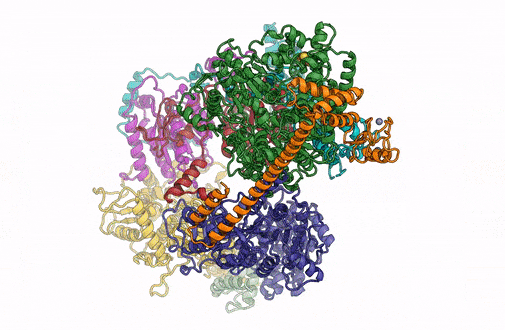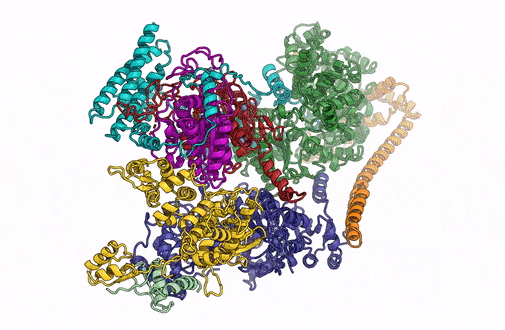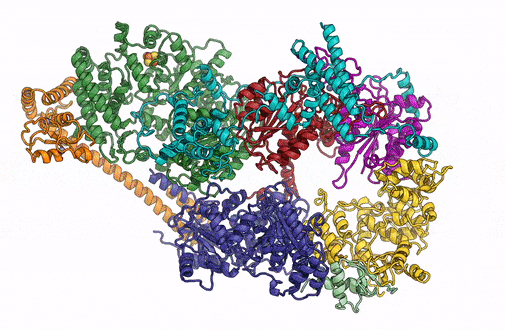
A 3D rendering of the overall architecture of the TFIIH complex. (Credit: Basil Greber/Berkeley Lab)
“When we solve these complex structures, we not only improve our understanding of how they work, but we critically update the way we think about them and are able to pose new questions and design future experiments,” said Basil Greber, a researcher in Berkeley Lab’s Molecular Biophysics and Integrative Bioimaging (MBIB) Division, a postdoctoral fellow at UC Berkeley, and first author on the study.
“Seeing which regions of the assembly, sometimes at the single amino acid level, are involved in different aspects of its function allows us to generate testable hypotheses about what happens to the molecules when something goes wrong, such as in the case of illness-causing mutations.”
The TFIIH molecular assembly is integral to the initiation of DNA repair and transcription – the process of copying DNA sequences into RNA, which is the first step of gene expression – in humans and all other complex life forms. To perform these tasks, the proteins must unwind the DNA double helix in a specific region and create a bubble of space where repair and transcription enzymes can interact with one of the strands. By examining cryo-EM maps, Greber and study leader Eva Nogales discovered new details about how eight of the ten proteins in the assembly physically support and regulate each other during these processes.
“Our detailed description of the human TFIIH structure now allows us to start to understand the functional consequences of these mutations,” said Nogales, a researcher in MBIB and head of the Bay Area Cryo-EM Facility (BACEM). “Getting here was a technical tour de force, as the assembly is very hard to purify from cells, it is hard to handle, and is highly flexible. This is the type of challenge my lab thrives on!”

First author Basil Greber (left) and study leader Eva Nogales (right) stand in front of their Titan Krios G2 cryo-electron microscope, one of the most powerful cryo-EM instruments currently available, at the Bay Area Cryo-EM (BACEM) facility in Berkeley, California. (Credit: Basil Greber)
Ever since the first 3D atomic map of a molecule was generated 29 years ago, cryo-EM technology has been a revolutionary tool for the fields of structural biology and biochemistry. Scientists based at Berkeley Lab contributed significantly to each stage of cryo-EM’s development, from helping establish the necessary early fundamentals of the technology in the 1970s, to enabling previously unprecedented resolution with the creation and subsequent commercial introduction of direct electron “counting” cameras in 2004. The BACEM microscope is equipped with the latest version of a direct detector camera.
This work was funded by the National Institute of General Medical Sciences, the Swiss National Science Foundation, and the Howard Hughes Medical Institute. The authors utilized resources of the National Energy Research Scientific Computing Center (NERSC), a Department of Energy Office of Science user facility at Berkeley Lab.
# # #
Founded in 1931 on the belief that the biggest scientific challenges are best addressed by teams, Lawrence Berkeley National Laboratory and its scientists have been recognized with 13 Nobel Prizes. Today, Berkeley Lab researchers develop sustainable energy and environmental solutions, create useful new materials, advance the frontiers of computing, and probe the mysteries of life, matter, and the universe. Scientists from around the world rely on the Lab’s facilities for their own discovery science. Berkeley Lab is a multiprogram national laboratory, managed by the University of California for the U.S. Department of Energy’s Office of Science.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.