Tag: microbes
Unlocking the Biochemical Treasure Chest Within Microbes

Decoding Messages in the Body’s Microscopic Metropolises
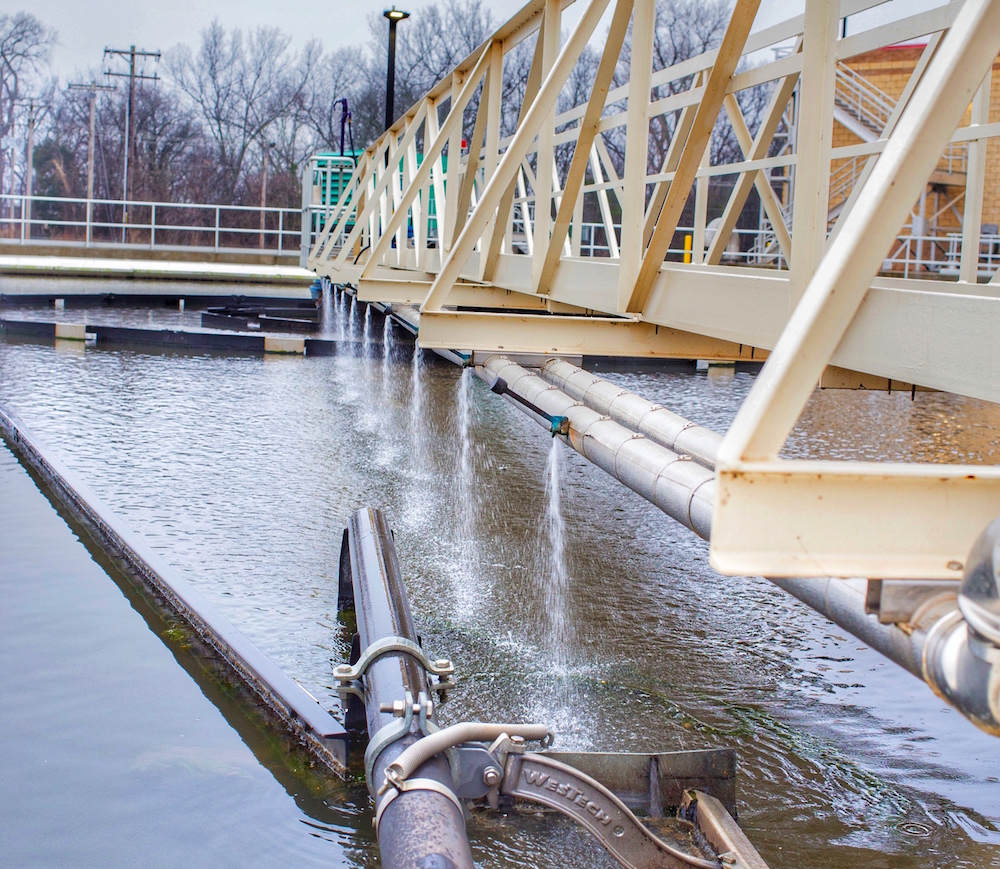
Can We Reuse Polluted Water? Yes, Add Bacteria

Scientists Hit Pay Dirt with New Microbial Research Technique
Berkeley Lab Technology Provides Clarity Amid Hawaiian Water Contamination Concerns

Understanding Microbiomes for Advanced Wastewater Treatment and Reuse Systems

Plants Really Do Feed Their Friends
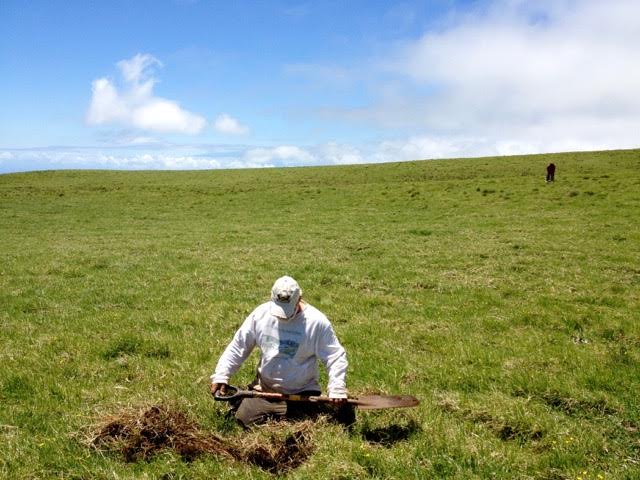
Digging Deep: Harnessing the Power of Soil Microbes for More Sustainable Farming
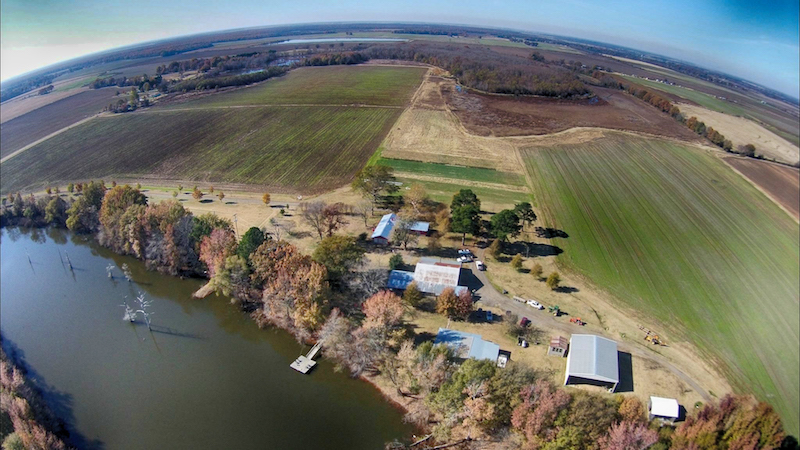

To Find New Biofuel Enzymes, It Can Take a Microbial Village

Berkeley Lab Researchers Help Map the Microbiome of Everything

What’s On Your Skin? Archaea, That’s What

Genes, Early Environment Sculpt the Gut Microbiome
