Tag: Advanced Light Source
Advanced Light Source Upgrade Approved to Start Construction

How X-Rays Can Make Better Batteries

Safely Studying Dangerous Infections Just Got a Lot Easier
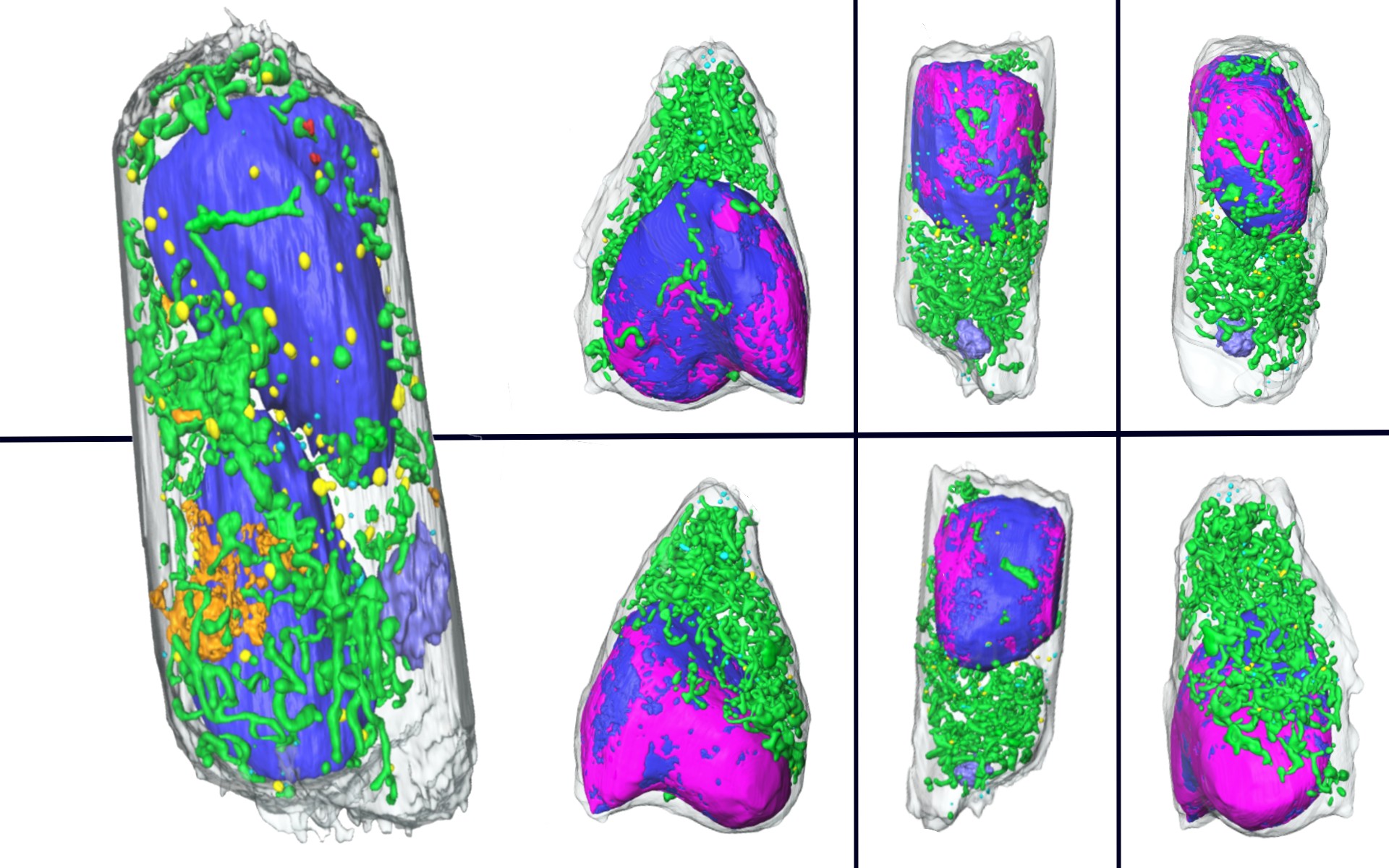
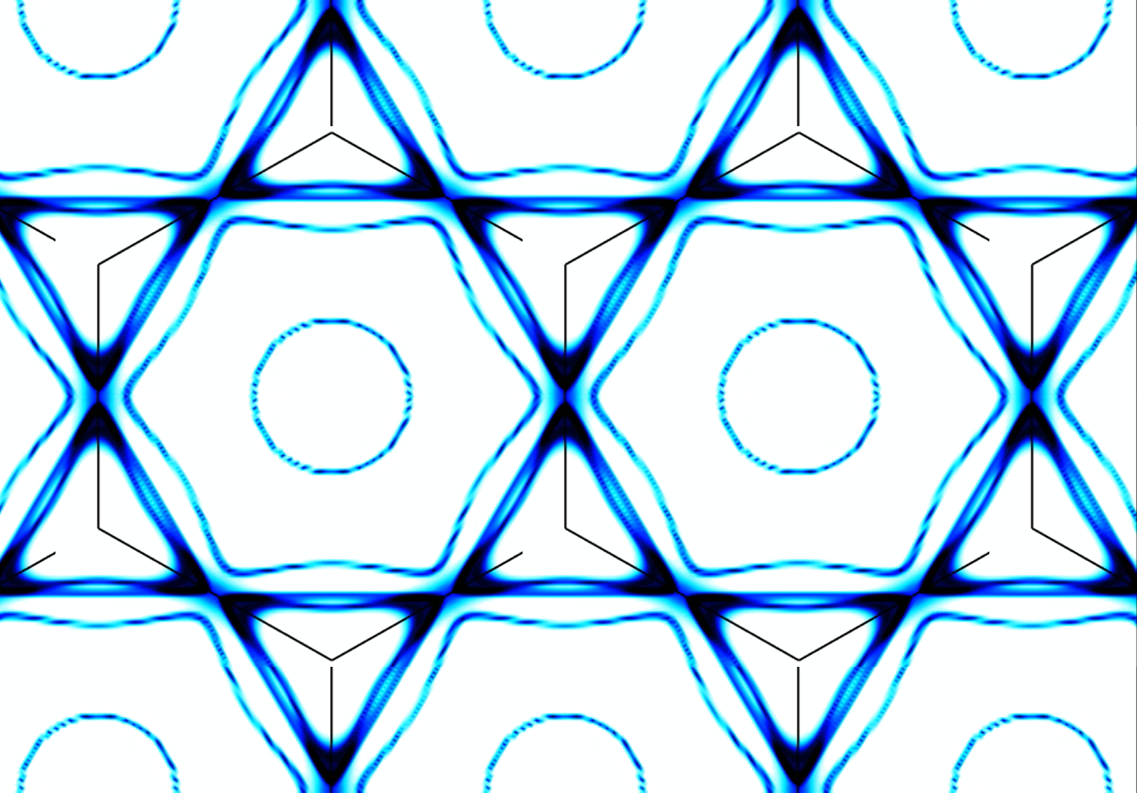
Scientists Discover ‘Secret Sauce’ Behind Exotic Properties of New Quantum Material

With a Little Help, New Optical Material Assembles Itself

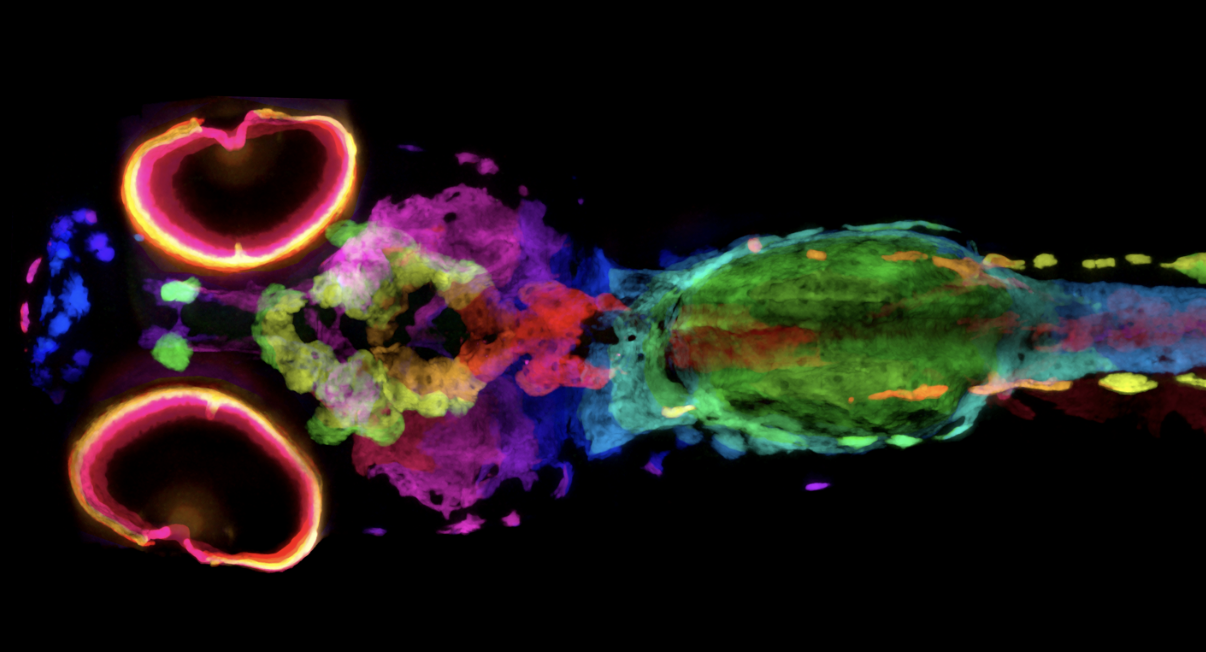
New Technique Visualizes Every Pigment Cell of Zebrafish in 3D
New Device Advances Commercial Viability of Solar Fuels

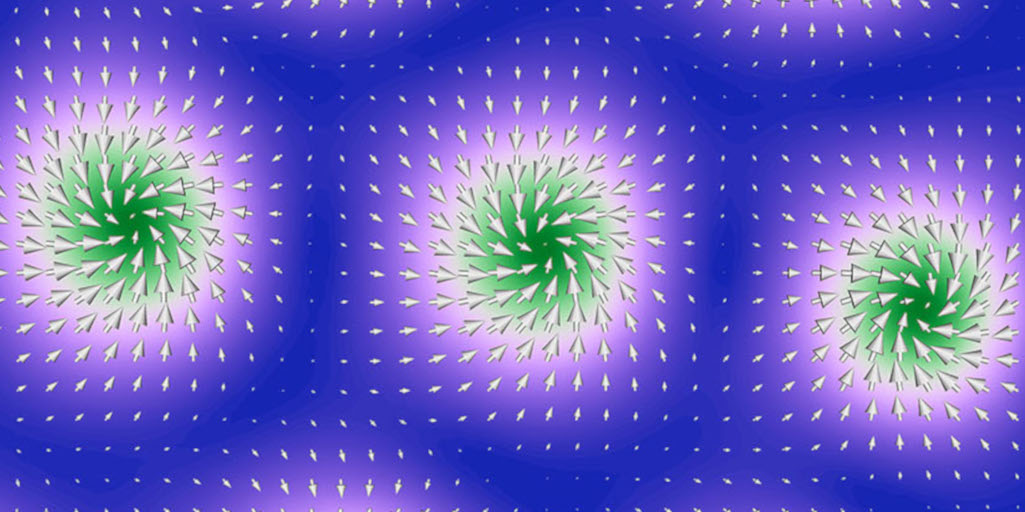
Skyrmions Could Be Future of Computing; X-Ray Experiments Reveal Their Secrets

New Technique Paves the Way for Perfect Perovskites

Cell ‘Fingerprinting’ Could Yield Long-Awaited Alzheimer’s Disease Diagnostic

Energy Secretary Jennifer Granholm visits Berkeley Lab
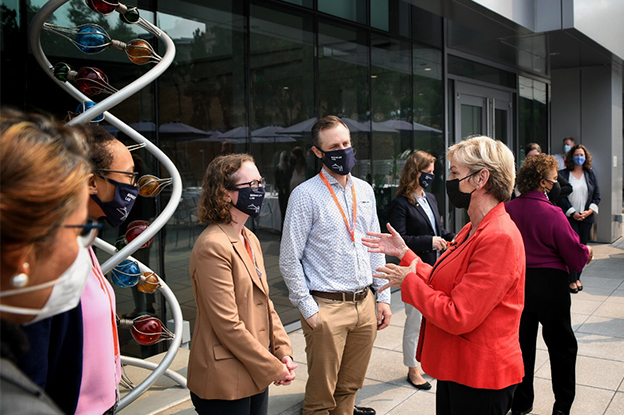

This Exotic Particle Had an Out-of-Body Experience; These Scientists Took a Picture of It
